Background
Output of Integrated Reaction Rates (IRR) is available in WRF-Chem from V4.0 and IRR can be analyzed with PERMM Analysis Tool. You can check the instruction on the UCAR website.
This post is the part2 which focuses on the application of PERMM for the WRF-Chem IRR data*.
If you wanna learn the basic usage of PERMM, please check part1.
If you wanna define your own mechanism file, please check part3.
Reactions related to O3
In the WRF-Chem, the reactions are written by the KPP and the IRR outputs are saved as IRR_DIAG_d0*_*** if you enable it.
We can print the reactions producing O3 using the PERMM without the IRR* files.
I will focus on the MOZART mechanism:
$ python -m permm -i --mechanism mozart_wrfchem
# all reaction names that create O3
>>> find_rxns(products=O3)
['CH3CO3_HO2_IRR', 'ISOP_O3_IRR', 'MACR_O3_IRR', 'MCO3_HO2_IRR', 'MVK_O3_IRR', 'NO3_HV_IRR', 'O_M_IRR']
# all reaction names related to O3 (either as reactants or products)
>>> find_rxns(reactants=O3, products=O3, logical_and=False)
['C10H16_O3_IRR', 'C2H4_O3_IRR', 'C3H6_O3_IRR', 'CH3CO3_HO2_IRR', 'HO2_O3_IRR', 'ISOP_O3_IRR', 'MACR_O3_IRR', 'MCO3_HO2_IRR', 'MVK_O3_IRR', 'NO3_HV_IRR', 'O3_HV_IRR', 'O3_HV_a_IRR', 'O3_NO2_IRR', 'O3_NO_IRR', 'OH_O3_IRR', 'O_M_IRR', 'O_O3_IRR']
# print all reactions that create O3
>>> print_rxns(products=O3)
ISOP_O3_IRR 1.0*ISOP + 1.0*O3 ->[k] 0.6*CH2O + 0.4*MACR + 0.3*CO + 0.27*OH + 0.2*MVK + 0.2*MCO3 + 0.2*CH3COOH + 0.1*O3 + 0.07*C3H6 + 0.06*HO2
MACR_O3_IRR 1.0*MACR + 1.0*O3 ->[k] 0.8*CH3COCHO + 0.7*CH2O + 0.275*HO2 + 0.215*OH + 0.2*O3 + 0.2*CO
MVK_O3_IRR 1.0*MVK + 1.0*O3 ->[k] 0.95*CH3COCHO + 0.8*CH2O + 0.2*O3 + 0.08*OH + 0.06*HO2 + 0.05*CO + 0.04*CH3CHO
CH3CO3_HO2_IRR 1.0*CH3CO3 + 1.0*HO2 ->[k] 0.75*CH3COOOH + 0.25*O3 + 0.25*CH3COOH
MCO3_HO2_IRR 1.0*HO2 + 1.0*MCO3 ->[k] 0.75*CH3COOOH + 0.25*O3 + 0.25*CH3COOH
NO3_HV_IRR 1.0*NO3 ->[j] 0.89*O3 + 0.89*NO2
O_M_IRR 1.0*M + 1.0*O ->[k] 1.0*O3
# print net reaction
>>> print_net_rxn(products=O3)
3*O3 + 2*HO2 + 1*CH3CO3 + 1*ISOP + 1*M + 1*MACR + 1*MCO3 + 1*MVK + 1*NO3 + 1*O ->[n] 2.89*O3 + 2.1*CH2O + 1.75*CH3COCHO + 1.5*CH3COOOH + 0.89*NO2 + 0.7*CH3COOH + 0.565*OH + 0.55*CO + 0.4*MACR + 0.395*HO2 + 0.2*MVK + 0.2*MCO3 + 0.07*C3H6 + 0.04*CH3CHO
IRRs
Combine IRR files
We need to calculate the regional mean IRRs based on each IRR file.
The script below simplifies the plot process laster and make it reproducing.
import os
import pandas as pd
import xarray as xr
import numpy as np
from glob import glob
from wrf import getvar
irr_dir = '/WORK/nuist_chenq_2/xin/data/history/NJ/chem/20190725/lfr_lnox_waccm25_aeroff/'
irr_file = sorted(glob(irr_dir+'irr/IRR_DIAG_d03_2019-07-25_0[3-6]*'))
wrfout_file = irr_file[0].replace('irr', 'wrfout').replace('IRR_DIAG', 'wrfout')
reactions = ['C10H16_O3_IRR', 'C2H4_O3_IRR', 'C3H6_O3_IRR', 'CH3CO3_HO2_IRR', 'HO2_O3_IRR', 'ISOP_O3_IRR', 'MACR_O3_IRR', 'MCO3_HO2_IRR', 'MVK_O3_IRR', 'NO3_HV_IRR', 'O3_HV_IRR', 'O3_HV_a_IRR', 'O3_NO2_IRR', 'O3_NO_IRR', 'OH_O3_IRR', 'O_M_IRR', 'O_O3_IRR']
# set the region and height range
crop_region = [118.98, 119.13, 31.9, 32]
bottom = 0 # km
top = 1 # km
# open irr and wrfout data
ds_irr = xr.open_mfdataset(irr_file, combine='nested', concat_dim='Time')
ds_wrfout = xr.open_dataset(wrfout_file)
def mean_irr(ds_wrfout, ds_irr, reactions):
'''Get the mean IRR in the crop_region and from bottom to top'''
# pick interested reactions
ds_irr = ds_irr[reactions]
# get basic info from the wrfout file
lon = ds_wrfout.XLONG.isel(Time=0)
lat = ds_wrfout.XLAT.isel(Time=0)
z = getvar(ds_wrfout._file_obj.ds, 'z', units='km')
# calculate the mean IRR in the region
subset = (lon>=crop_region[0]) & (lon<=crop_region[1]) & (lat>=crop_region[2]) & (lat<=crop_region[3]) \
& (z >= bottom) & (z <= top)
# we use the "difference" here as the IRRs are integrated values
ds_irr = ds_irr.where(subset.drop_vars(['Time', 'XTIME']), drop=True)\
.mean(dim=['south_north', 'west_east', 'bottom_top'])\
.diff('Time')
return ds_irr
def conv_da(ds_irr):
'''Convert IRR Dataset to DataArray and add coords'''
# get the datetime for the Time coordinate
times = pd.to_datetime(pd.Series([os.path.basename(f) for f in irr_file]), format='IRR_DIAG_d03_%Y-%m-%d_%H:%M:%S')[1:]
times = times.to_xarray().drop_vars('index').rename({'index': 'Time'})
# set the time coords and rename
da_irr = ds_irr.assign_coords({'Time': times}).to_array(dim='RXN', name='IRR').transpose()
# convert the unit to pptv/s
delta = times.diff(dim='Time')[0] / np.timedelta64(1, 's') # seconds
da_irr = da_irr * 1e6 / delta.values
da_irr.attrs['units'] = 'pptv/s'
return da_irr
ds_irr = mean_irr(ds_wrfout, ds_irr, reactions)
da_irr = conv_da(ds_irr)
# save to netcdf
comp = dict(zlib=True, complevel=9)
da_irr.to_netcdf('test.wrfchem.nc', encoding = {'IRR': comp}, compute=True, engine='netcdf4')
What you need to modify is irr_dir and irr_file.
Of course, if you wanna calculate the mean IRRs based on the bottom_up levels, then we don’t need the wrfout files anymore.
Anyway, I get the z (km) from the wrfout file using wrf-python.
When the script is finished successfully, you will get the nc file called test.wrfchem.nc.
Data structure
This is the overview of combined IRR netcdf file byxarray:
>>> import xarray as xr
>>> ds = xr.open_dataset('test.wrfchem.nc')
>>> ds
<xarray.Dataset>
Dimensions: (RXN: 17, Time: 23)
Coordinates:
* Time (Time) datetime64[ns] 2019-07-25T03:10:00 ... 2019-07-25T06:50:00
* RXN (RXN) object 'C10H16_O3_IRR' 'C2H4_O3_IRR' ... 'O_M_IRR' 'O_O3_IRR'
Data variables:
IRR (Time, RXN) float32 ...
The IRR is the only data variable with the Time and RXN dimension.
As the coordinates have been added before, we can access them easily to plot the IRR.
Plot IRRs
As we have written the plot function in the part1 tutorial, we can edit it for the WRF-Chem IRR files this time.
python -W ignore -m permm --mechanism mozart_wrfchem test.wrfchem.nc --scripts plot_irr_wrfchem.py
Here’s the content of the script called plot_irr_wrfchem.py:
import xarray as xr
import proplot as plot
import numpy as np
# get the basic info of reactions
reaction_names = find_rxns(reactants=O3, products=O3, logical_and=False)
reaction_eqs = get_rxns(reactants=O3, products=O3, logical_and=False)
reaction_dict = dict(zip(reaction_names, [eq.display(digits = None) for eq in reaction_eqs]))
reactant_names = find_rxns(reactants=O3)
# read the reaction file
ds = xr.open_dataset('./test.wrfchem.nc')
da = ds.sel(RXN=reaction_names)['IRR']
# multiple the irr by net coefficient
coefficient = [eq['O3'].base.item() for eq in reaction_eqs]
da *= coefficient
# we only want to plot the large contributions
# so, sorting the integrated IRR by absolute vlalues and picking the 8 of them for plot
large_contrib = da.sum(dim='Time').sortby(abs(da).sum(dim='Time'), ascending=False).RXN[:8].values
irr_subset = da.sel(RXN=large_contrib)
# sort again for plot by real values
irr_subset = irr_subset.sortby(irr_subset.sum(dim='Time'))
# generate labels based on RXN_**
labels = []
for rxn in irr_subset.RXN.values:
labels.append(reaction_dict[rxn])
# plot
fig, axs = plot.subplots()
cmap = plot.Colormap('viridis')
lines = []
for i in range(irr_subset.sizes['RXN']):
# "step": https://matplotlib.org/3.1.1/gallery/lines_bars_and_markers/step_demo.html#sphx-glr-gallery-lines-bars-and-markers-step-demo-py
line = axs.step(irr_subset.Time,
irr_subset.isel(RXN=i),
where='pre',
label=labels[i],
color=cmap(i / (len(irr_subset.RXN)-1)),
)
lines.append(line)
# plot the total IRR
twin_ax = axs.alty(ycolor='gray6')
total_irr = da.sum(dim='RXN').rename('Total IRR')
total_irr.attrs['units'] = da.attrs['units']
total_line = twin_ax.step(total_irr, where='pre', label='Total IRR', color='gray6')
twin_ax.format(grid=False)
# set axis
axs.format(xlabel='Time (UTC)',
ylabel=f'IRR ({da.attrs["units"]})',
xlim=(irr_subset.Time.values[0], irr_subset.Time.values[-1]),
xlocator=('minute', range(0, 60, 30)),
xminorlocator=('minute', range(0, 60, 10)),
xformatter='%H:%M',
title='Mean IRRs between the surface and 1 km',
grid=False,
)
fig.legend(lines.append(total_line), loc='b', ncols=2)
fig.savefig('irr_o3_wrfchem.png')
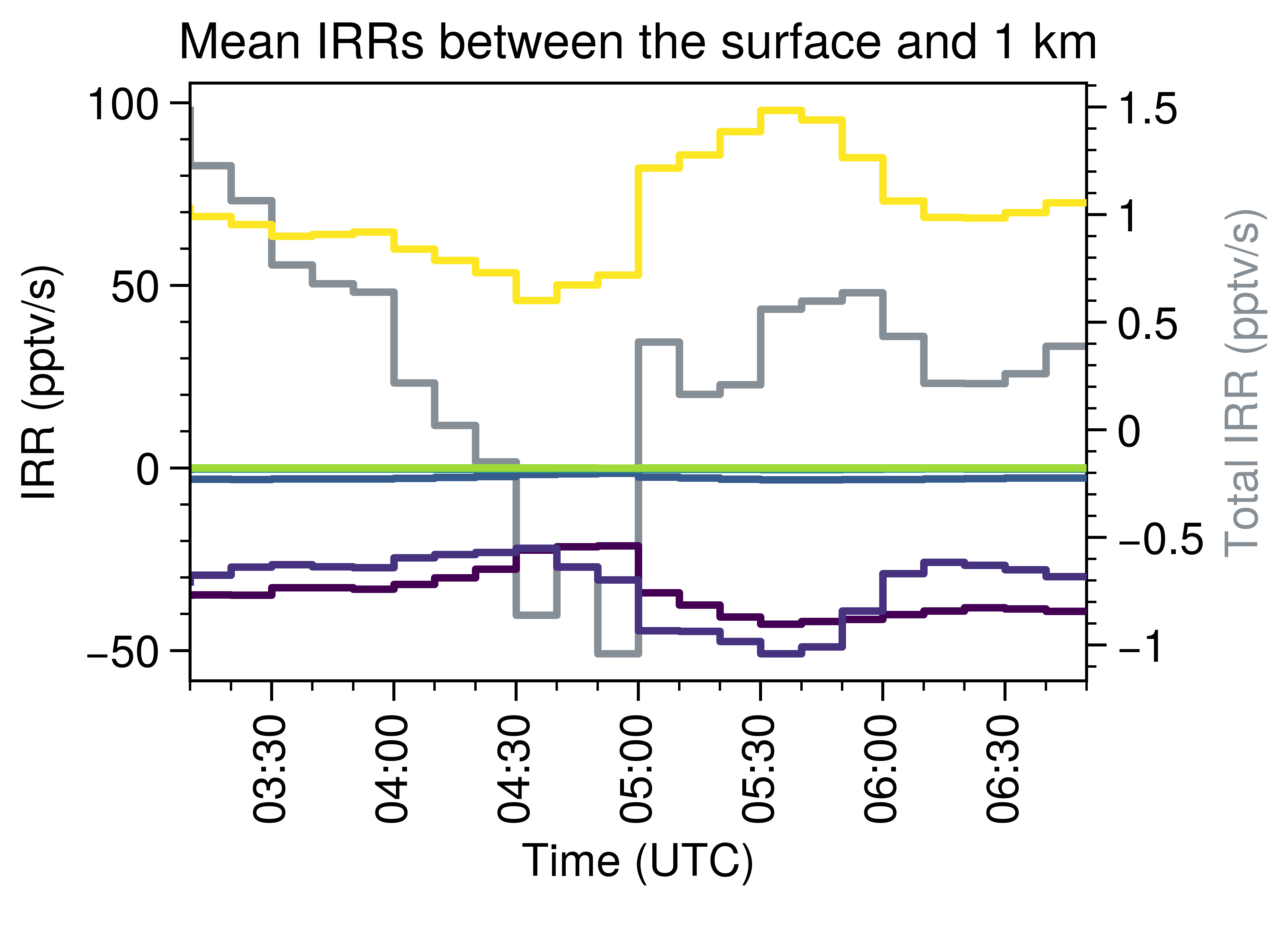
Here’s the result:
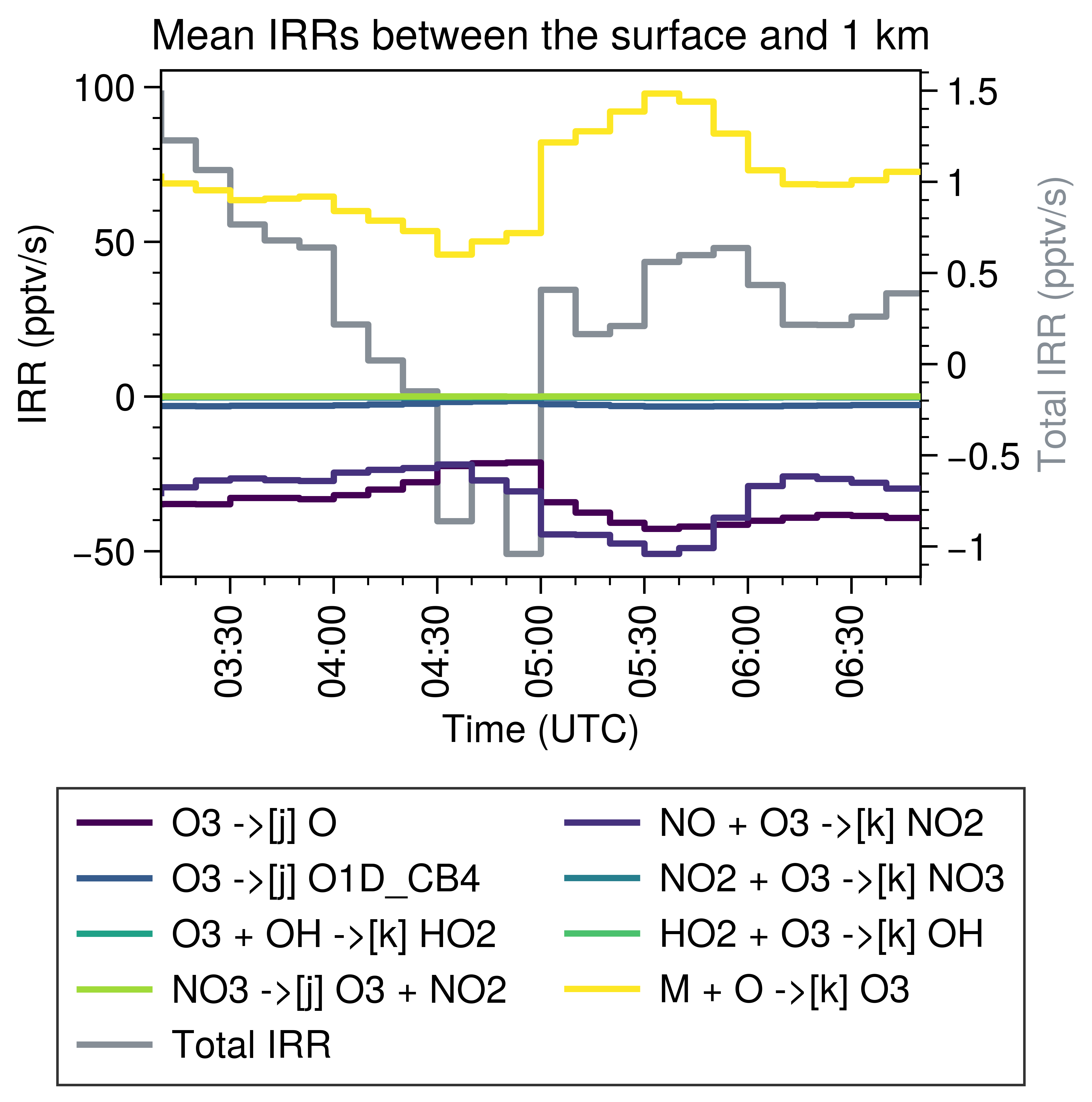
Note that the simulation is for one convection case, so the decrease of O3 at the noon is reasonable.
Version control
| Version | Action | Time |
|---|---|---|
| 1.0 | Init | 2020-12-01 |
| 1.1 | Update coefficients | 2020-12-02 |
Say something
Thank you
Your comment has been submitted and will be published once it has been approved.
OOPS!
Your comment has not been submitted. Please go back and try again. Thank You!
If this error persists, please open an issue by clicking here.